COVID-19: An unpredictable winter marked by the massive and unprecedented spread of the Omicron variant
Mircea T. Sofonea, University of Montpellier and Samuel Alizon, French National Center for Scientific Research (CNRS)
A certain degree of “pandemic fatigue” has set in among part of the population, but SARS-CoV-2 continues to evolve. As France experiences its eighth wave (the fourth of 2022), dominated by the Omicron BA.5 subvariant, another subvariant, known as BQ.1.1, is spreading rapidly. Samuel Alizon, research director (CNRS, CIRB), and Mircea Sofonea, associate professor (University of Montpellier, MIVEGEC), discuss the health situation expected this winter and highlight the challenges facing surveillance and research in our country. What are the implications?

The Conversation: With the end of summer, the news has once again turned to COVID-19 and its variants. Now it’s BQ.1.1 and other XBB variants that are being discussed. What can we already say about these “subvariants”? And how are they being monitored?
Samuel Alizon: Since September, we have seen significant diversification of SARS-CoV-2, with the emergence of numerous subvariants of the Omicron variant.
BA.4.6, BA.2.75, BA.5.2, and even B.1.1.529.5.3.1.1.1.1.1.1, renamed BQ.1.1 according to the Pango nomenclature, which provides an identification system for tracking SARS-CoV-2 lineages of epidemiological interest…
All of these lineages, which are prevalent in various regions of the world (BQ.1.1, for example, is currently spreading rapidly in France), are officially classified as part of the Omicron variant. They are therefore subvariants, but in reality, they could easily be described as variants.
TC: What do we know about these new subvariants? Do they pose an epidemiological threat?
SA: At this stage, much of what is known about these new strains should be taken with a grain of salt, as the information comes, at best, from preprints that have not undergone peer review. Little is known about their virulence and, of course, virtually nothing about the long-term effects of infection.
We are certain, however, of the mutations present in the genomes of these lineages, since they are what define them. The BA.5 variant, for example, had established a mutation at position 452 of the spike protein. This mutation was the subject of extensive study, as it was characteristic of the Delta variant at the time of its emergence.
For BQ.1.1, a completely different set of mutations is observed in the Receptor Binding Domain (RBD) of this protein—in other words, the part of the spike protein that interacts with ACE2, the “lock” located on the surface of the cells infected by SARS-CoV-2. This is particularly true of the S:R346T mutation. As indicated in a preprint from a consortium of Chinese research teams, these mutations appear to confer significant immune evasion potential to this lineage. Furthermore, BQ.1.1 may not be susceptible to treatments with monoclonal antibodies available in France (such as the tixagevimab-cilgavimab combination (Evusheld)).
Among the subvariants being monitored—particularly due to their potential ability to evade the immune system—is the XBB lineage, which arose from a recombination between viruses of the BJ.1 and BM.1.1 lineages during co-infection of a single cell.
TC: How are these variants of concern monitored? Where does the epidemiological data come from?
SA: From an epidemiological standpoint, the quality of the UK’s surveillance system remains outstanding. Their latest report, dated October 7, 2022, which combines testing and sequencing data, provides a particularly clear picture of the current epidemiological situation.
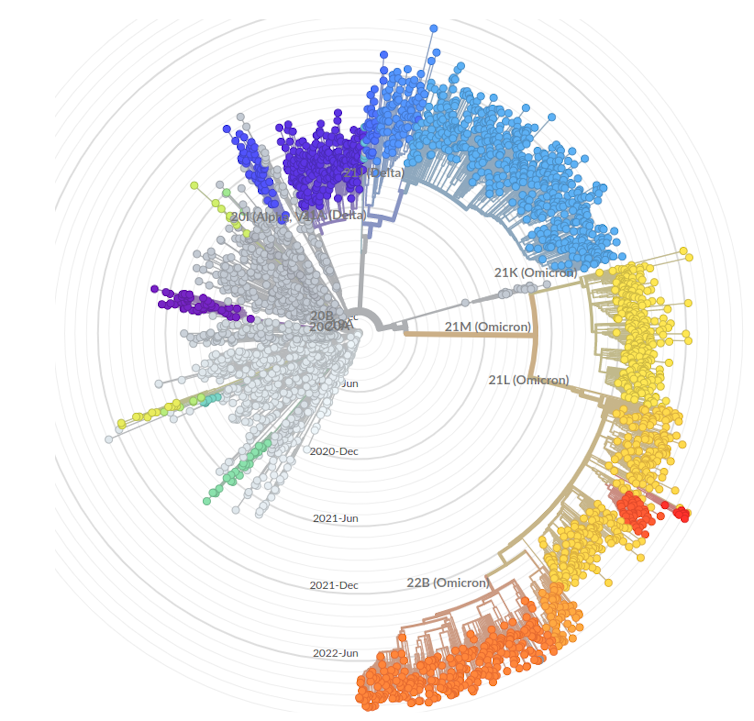
For other countries, including France, we rely on sequencing data shared on the GISAID platform. Several websites, including Nextstrain.org and the excellent covSPECTRUM developed by Prof. Tanja Stadler’s team in Switzerland, allow users to visualize the dynamics of variants in real time (within the limits of the data provided by each country).
Mircea Sofonea: It should be noted that, unlike previous waves caused by the emergence of a new variant, screening data from RT-qPCR tests (which detect predefined mutations and are therefore used to track already known variants) no longer allow us to distinguish these new lineages.
This deprives us of an early—and therefore valuable—indicator for feeding real-time data on current replacement dynamics into the models. These dynamics can then only be determined through sequencing, with a delay of at least one week after sample collection (which itself occurs several days after the onset of infection) and based on a sample size that, for logistical reasons, is limited – the Flash surveys conducted by the EMERGEN consortium, covering 1,000 to 2,000 interpretable sequences.
The problem is that, this time, France is the first country to see the new (sub)variant (BQ.1.1) become dominant. So we can no longer rely on the trends observed across the Channel!
As the genetic diversity of SARS-CoV-2 once again puts our surveillance and healthcare systems to the test, the technical and scientific infrastructure—from individual testing to population-level analysis—upon which our collective preparedness relies requires immediate investment commensurate with the public health stakes. https://cov-spectrum.org/embed/VariantInternationalComparisonChart?json=%7B%22variantSelector%22%3A%7B%22location%22%3A%7B%7D%2C%22dateRange%22%3A%7B%22mode%22%3A%22Past6M%22%7D%2C%22variant%22%3A%7B%22nextcladePangoLineage%22%3A%22bq.1.1*%22%7D%2C%22samplingStrategy%22%3A%22AllSamples%22%7D%2C%22countries%22%3A%5B%22France%22%5D%2C%22logScale%22%3Afalse%7D&sharedWidgetJson=%7B%22originalPageUrl%22%3A%22https%3A%2F%2Fcov-spectrum.org%2Fexplore%2FFrance%2FAllSamples%2FPast6M%2Fvariants%3FnextcladePangoLineage%3Dbq.1.1*%26%22%7D
TC: How do these variants fit into the overall trend observed for Omicron?
SA: Up until then, we often saw a dominant lineage and fairly pronounced shifts between variants: Alpha replaced all the lineages that came before it, then Delta replaced Alpha, and so on.
As can be seen on the Nexstrain.org website, the sublineages derived from BA.2, BA.4, and BA.5 appear to be circulating simultaneously worldwide.
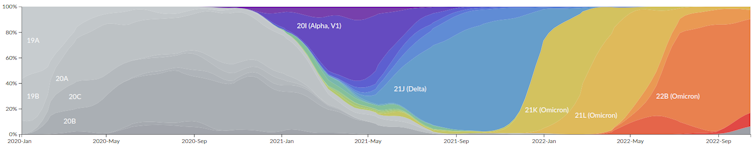
As for the reasons behind this diversification, it is obviously impossible to be certain. The study of biological diversity is a field in its own right, but we can put forward two hypotheses.
On the one hand, the diversity of parasites is often correlated with that of their hosts. However, human populations are now more diverse than ever in terms of their immunity, whether acquired through vaccination or following infection.
On the other hand, the spread of the virus is also proportional to the number of new infections, and currently, in many countries, the virus is spreading unchecked.
[More than 80,000 readers rely on The Conversation’s newsletter to better understand the world’s major issues. Subscribe today]
TC: Could we have anticipated such a situation?
MS: While it is currently impossible—and will likely remain so for many years to come—to predict the exact evolutionary trajectory of a coronavirus, the diversification trends observed since the spring are not necessarily surprising.
The first two years of the pandemic were marked by strong selection pressures favoring transmissibility—the Alpha and Delta variants were able to emerge only because their particularly high transmissibility allowed them to overcome the effects of social distancing and mask-wearing.
Omicron, which originated from an older branch of the SARS-CoV-2 phylogenetic tree rather than from those two variants, spread in a context where population immunity (from prior infection and vaccination) had become an additional barrier to viral transmission. Since the spring of 2022, this immunity has, in fact, been the only remaining barrier.
Thanks to its combination of high transmissibility and strong immune evasion, Omicron BA.1 has virtually driven other lineages to extinction and has been able to circulate largely unchecked, with little to no resistance from the humoral and cellular immunity developed against them.
However, since every infection gives rise to (random) mutations, the conditions were ripe for the establishment of a “conveyor belt” of diversification fueled by immune escape.
TC: So is this capacity for diversification limitless?
SA: This question prompts us to revisit the discussions from 2020 and even 2021, when some predicted that the virus would “run out of steam” and that its ability to mutate would gradually diminish.
In fact,viral evolution is always difficult to predict because each mutation can completely alter the virus’s “adaptive landscape”—that is, its potential for evolution. In addition to genetic constraints, there are also environmental factors and the variability of the populations infected by the virus.
An interesting aspect of the current diversification of viral lineages is that there are a number of cases of parallel evolution, that is, lineages that independently acquire the same mutation.
TC: What might the potential consequences be for this winter? Are forecasts still possible?
SA: It’s very difficult, because beyond the scientific challenges, research teams in France have virtually no annual core funding (known as “recurrent” funding) left, and many projects have been rejected this year. In short, we are no longer able to explore prospective scenarios. And unlike in 2020 and 2021, there is no longer a scientific advisory body to request such analyses. The uncertainties regarding how the winter will unfold are therefore enormous.
What can be said, however, is that the current wave (which began in mid-September) is still dominated by the BA.5 variant, which had already caused the third surge of the pandemic in 2022, back in July. This fourth wave likely has multiple causes, but it is worth noting that it coincided with the start of the school year and was first observed circulating among children.
As for why the same variant was able to trigger a new wave so soon after the previous one, it is likely that social factors played a decisive role. The summer likely brought the previous wave to an end, while the start of the school year likely contributed to the resurgence of viral transmission. The persistence of post-infection immunity from the summer wave for BA.5 likely explains why the peak was reached so quickly among children (before the end of September).
For adults, the outbreak naturally began later, following the spread of the outbreak among children. Unfortunately, the peak of the outbreak has been slow to emerge this time. Sequencing data are still incomplete, but as suggested by the visualizations on the covSPECTRUM website, the rise of the “new” variants mentioned above (such as BQ.1.1) may be the cause.
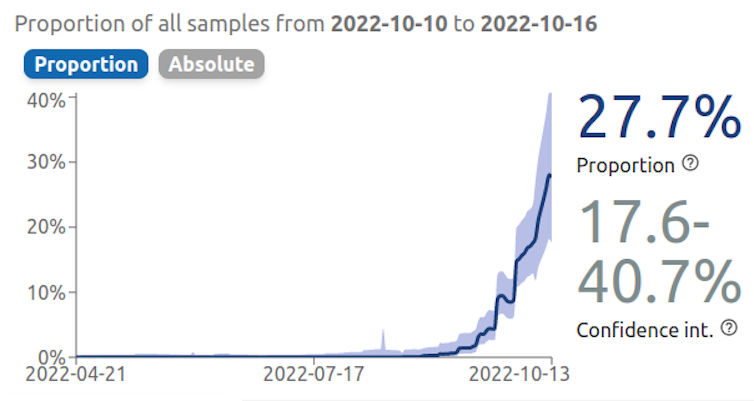
If this is confirmed, there is concern that a fifth wave in 2022 could overlap with the fourth, although school holidays should help ease the situation. In the absence of a national vaccination campaign, the severity of this wave would be directly linked to immune escape or, more precisely, to the extent of cross-immunity between BA.5 and new variants.
MS: Short-term forecasts (i.e., over a two-week horizon) are still possible by taking into account the epidemic’s momentum and the history of past waves. The current eighth wave has so far been in line with previous waves in terms of intensity and duration.
Monthly projections, however, require certain assumptions to be made, particularly regarding the kinetics of post-infection and post-vaccination immune decline, which are being reshaped by new subvariants… These assumptions cannot be properly informed at this time, while inferences based on hospital and testing data are increasingly difficult to draw, and, unfortunately, there is a lack of support for developing more automated and robust approaches.
TC: Winter is also the peak season for other outbreaks… Are there any common insights regarding flu and COVID-19 outbreaks?
SA: Yet another issue that makes us regret the lack of support for epidemiological modeling teams in France! Yes, the co-circulation of influenza and SARS-CoV-2 is a cause for concern this year because early data from the Southern Hemisphere suggest that, unlike in 2021 and, especially, 2020, we may see “normal” influenza circulation.
There are two potential consequences to be concerned about: some direct, as co-infection with both viruses may be more severe, and others indirect, as hospital capacity may become overwhelmed more quickly.
MS: The flu began circulating again last year, and the previous season was particularly unusual, with a late peak in March and April, just as we were gradually easing up on our vigilance and protective measures regarding SARS-CoV-2 (wearing masks, handwashing, etc.)—measures that also prevent the spread of the flu virus.
Currently, flu activity indicators in mainland France (with Réunion and Martinique in the epidemic phase) are comparable to levels seen before the COVID-19 pandemic.
Betweenthe current bronchiolitis outbreak, which is affecting the entire mainland, and the upcoming flu season, a potential ninth wave of COVID-19 will further complicate the work of healthcare workers who have been strained by nearly three years of constant pressure.
There is an urgent need to strengthen our capacity for monitoring, predicting, and simultaneously controlling respiratory viral infections. The methodological challenges posed by this public health issue are highly specific, and unfortunately, we currently lack the resources to address them.
While it is entirely legitimate—and desirable, particularly to avoid exacerbating “pandemic fatigue”—that daily attention should not be focused on COVID-19, it is nonetheless essential that, behind the scenes, we support this transition toward a controlled endemic circulation of the virus, rather than one that is simply endured as it is now. Research can help bring this goal closer to reality.
Mircea T. Sofonea, Associate Professor of Epidemiology and the Evolution of Infectious Diseases, MIVEGEC Laboratory, University of Montpellier and Samuel Alizon, Research Director Research CNRS and Director of the Ecology and Evolution of Health team at the Interdisciplinary Research Center in Biology (CIRB) UMR 7241 – U1050 Inserm, National Center for Scientific Research (CNRS)
This article is republished from The Conversation under a Creative Commons license. Readthe original article.