New structures identified within our chromosomes
Since the beginning of this century, major technological advances in molecular and genomic science, combined with new approaches in ultra-high-resolution photonic microscopy, have revealed new chromosomal structures and new principles governing the organization of our genomes (the genome contains all genetic information and thus all DNA sequences transmitted through cell division).
Frédéric Bantignies, University of Montpellier and Giacomo Cavalli, University of Montpellier
This knowledge is crucial not only for understanding how our cells function, but also for gaining a better understanding of certain diseases and their progression.
Deoxyribonucleic acid, or DNA, contains the genetic information specific to each living species. Chromosomes serve as the physical carriers of this information. The complete set of chromosomes is also called the genome and contains all the genes. Since the discovery in 1953 of DNA’s double-helix structure, the question of its organization within the nuclei of our cells has sparked extensive global research. To fully grasp this issue, it is worth noting here that humans have 46 chromosomes, which contain 3 billion base pairs (bp) of DNA, representing a molecular strand approximately 2 meters long. Yet this strand is capable of fitting inside cell nuclei that are about 10 μm (one micrometer = one-thousandth of a millimeter) in diameter. To better understand how the genome and our genes function, it was therefore essential to understand how all this DNA is able to fold up within a cell nucleus.
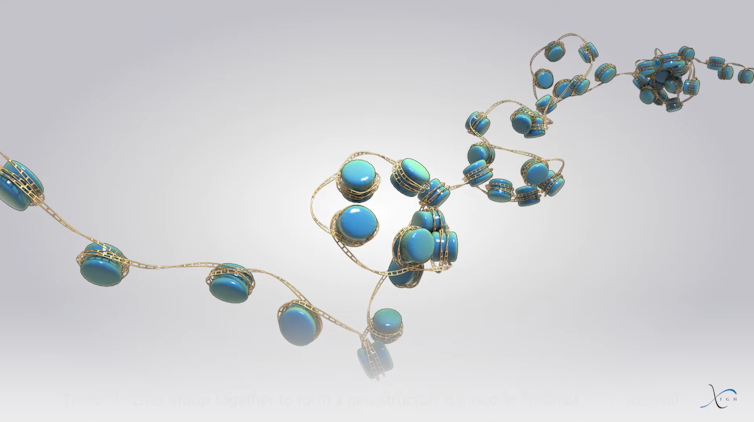
At the beginning of this structure is the nucleosome, whose structure was elucidated in 1997. The nucleosome is composed of highly basic proteins called histones, which have a strong affinity for the acidic DNA molecule. These histones form a central core around which the DNA molecule winds, at a rate of 146 base pairs per nucleosome, creating a sort of “cookie” 11 nm (one nanometer = one millionth of a millimeter) wide and 6 nm high. The nucleosome thus represents the first structural level of DNA organization within the nucleus. A sign of its importance is that this very distinctive structure is found in all organisms with nuclei (eukaryotes), whether they are unicellular or more complex, such as animals and plants.
For a very long time, it was believed that the sequence of nucleosomes (also known as chromatin fibers), forming a sort of “string of pearls,” wound itself in a regular pattern to form a fiber 30 nm in diameter. Spiral supercoils of this fiber were thought to ultimately form chromosomes. However, technical limitations—notably the low resolution of microscopy techniques and the lack of alternative methods—prevented scientists from verifying whether this hypothesis was correct.
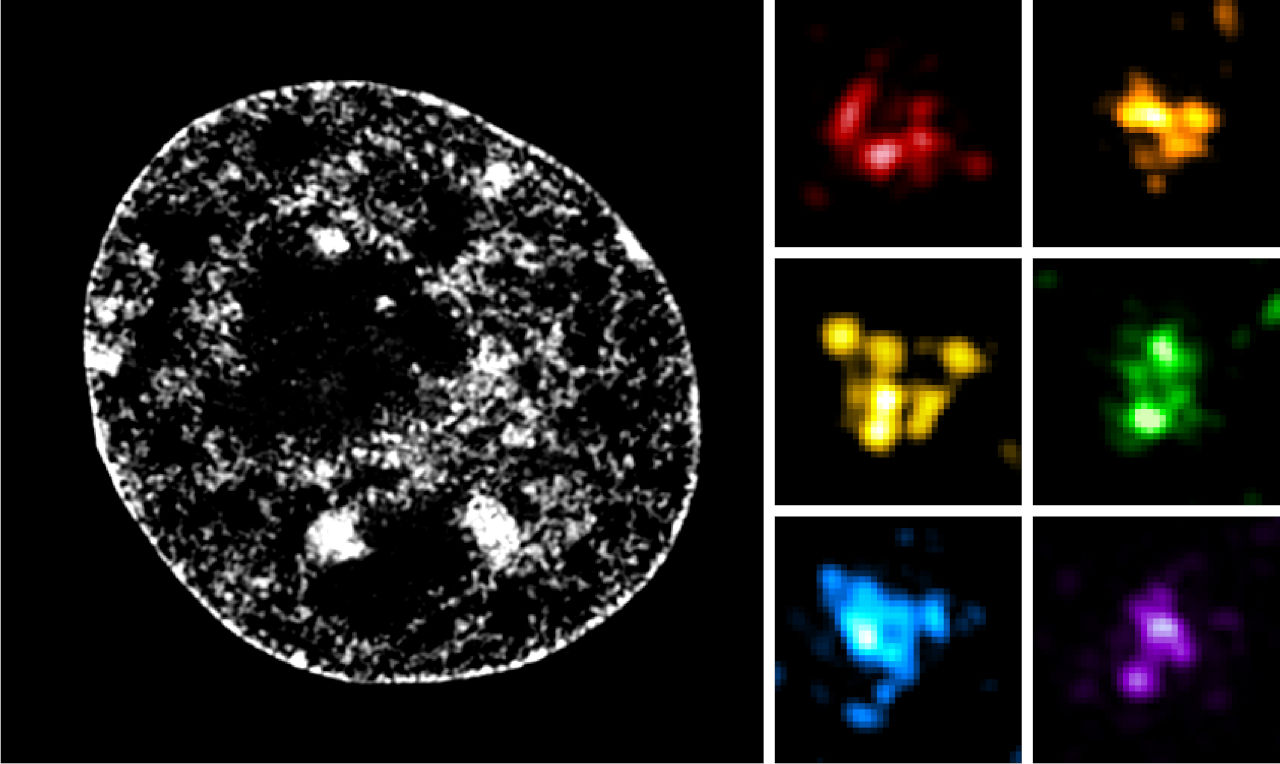
On the other hand, it has been known since 1985 that the organization of chromosomes within cell nuclei is not random. Furthermore, the well-known X-shaped form of chromosomes is not incorrect, but represents only a very transient stage of their organization. Indeed, this highly condensed X-shaped conformation facilitates their segregation (distribution) into daughter cells during cell division. But the rest of the time, the shape of chromosomes is quite different. Visualizing them using fluorescent molecules capable of specifically binding to the DNA double helix has shown that each chromosome occupies its own territory within the nucleus, thereby avoiding excessive entanglement with other chromosomes. This property of “chromosomal territory” is also found in most species and appears to be very important, particularly for species with a large number of chromosomes.
New technologies offer a fresh perspective on the organization of the genome
At the beginning of the 21st century, extensive research conducted around the world has led to a better understanding of the different levels of the genome’s structural organization, ranging from the nucleosome to chromosomal regions.
These advances were made possible by the development and use of entirely new technologies. First, there were new genomics techniques, including ultra-high-throughput DNA sequencing (next-generation sequencing) and the ability to capture the fine molecular structures of chromosomes using the “Hi-C” method. At the same time, the advent of super-resolution photonic microscopy, which uses fluorescent DNA markers, has made it possible to visualize these chromosomal structures directly within the cell nucleus.

Frédéric Bantignies, Giacomo Cavalli, Courtesy of the author
Let’s return to our scale of chromosome organization. Its first level is the nucleosome. A second level of organization corresponds to clusters of a few nucleosomes, like small clusters known as “nucleosome clutches” (they were named this way by the researchers who discovered them, drawing an analogy to eggs found in brooding nests). Nucleosomes are therefore not arranged in a regular pattern as previously thought, but rather in irregular clusters.
These "nucleosome clusters" then group together to form a structure called a "Chromatin Nanodomain, " or CND, which encompasses approximately 100,000 to 200,000 DNA base pairs, forming large, irregular clusters of nucleosomes 150 to 300 nm wide. These two levels were discovered recently (in 2015 and 2020, respectively), thanks to super-resolution microscopy, which is capable of resolving structures ranging from 20 to 100 nm.
TADs organize the genome and regulate gene expression
The next level of this organization is called a TAD, short for “Topologically Associating Domain,” which was identified in 2012 using the Hi-C molecular method. TADs are composed of several CNDs, forming superclusters of nucleosomes approximately 500 nm wide. They thus encompass DNA segments of varying sizes, with an average of approximately 1 megabase (1 million base pairs). Our laboratory, in fact, contributed to the discovery of CNDs and TADs.
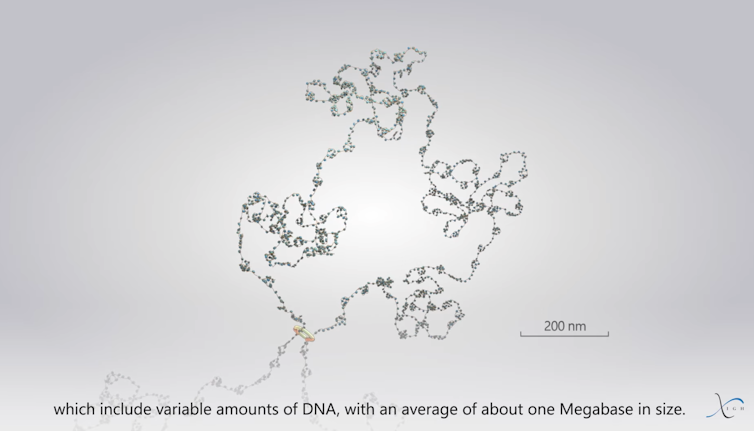
Frédéric Bantignies, Giacomo Cavalli, Courtesy of the author
TADs are fairly heterogeneous structures, largely due to their dynamic formation mechanism. This mechanism involves the passage of the famous “string of beads” (referring to the sequence of nucleosomes) through the ring formed by cohesin. The chromatin fiber continues to pass through the ring until it encounters nuclear factors called CTCF at the boundaries of a TAD; these act as a sort of “customs officer” that, stationed on the DNA, blocks the fiber’s progression. As the fiber passes through, the nucleosomes will organize themselves into clusters and CNDs. The TAD then represents the entire large chromatin loop that has passed through the cohesin ring.
Within chromosomes, gene activity is influenced by a wide variety of regulatory sequences (which act as switches) that can be located tens of thousands of base pairs away from their target gene. Thus, TADs keep genes and their regulatory regions within the same molecular environment, which can facilitate their expression (i.e., their transcription leading to protein production) in a specific cell type where their activity is required. Thanks to their boundaries, they also help keep genes well separated from one another, preventing active genes from influencing other inactive genes within a given cell type.
Recent studies have shown that chromosomal defects at the boundaries of TADs (such as inversions or deletions that affect the positioning of CTCF or render it nonfunctional) lead to defects in gene isolation and, consequently, to erroneous gene activation. In some cases, these rearrangements trigger the activation of genes known as “proto-oncogenes,” which can cause cellular transformation and lead to the development of tumors.
Compartmentalization and territorialization of the genome

Frédéric Bantignies, Giacomo Cavalli, Provided by the author
Several TADs then group into two distinct compartments: compartment “A,” which contains mainly active genes, and compartment “B,” which contains mainly inactive genes. This compartmentalization helps to enhance the functions of the genome.
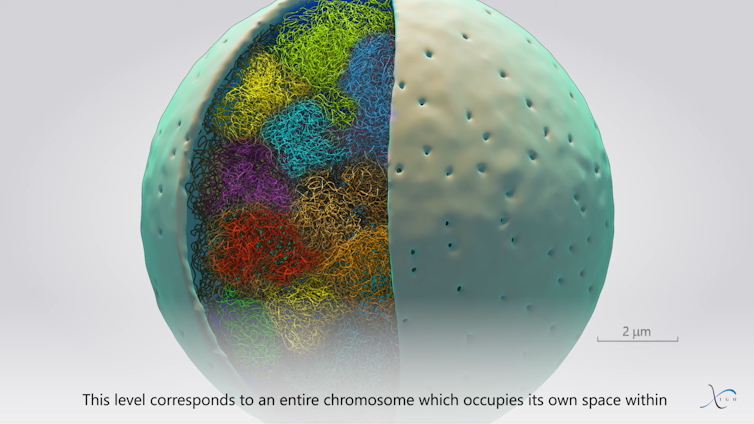
Frédéric Bantignies, Giacomo Cavalli, Courtesy of the author
Finally, at the end of this organizational scale are the chromosomal regions, which allow chromosomes to be distinguished from one another within the nucleus. This organization plays an important role during DNA replication (the exact copying of genetic material) and cell division, where each duplicated chromosome is split into two in the daughter cells.
And so we’ve come full circle! Well, almost: the next step is to better understand how this organization influences the various molecular processes inherent to the genome, which regulate, in particular, gene expression, DNA replication, DNA repair in the event of damage or stress, and DNA recombination, especially in reproductive cells. The field of investigation remains vast, but a better understanding of genome organization will pave the way for a deeper understanding of all these nuclear processes in normal cells—and, of course, in cells with chromosomal defects that lead to diseases such as cancer.![]()
Frédéric Bantignies, Director of Research , University of Montpellier and Giacomo Cavalli, CNRS Research Director, University of Montpellier
This article is republished from The Conversation under a Creative Commons license. Readthe original article.