Omicron, Delta, Alpha… Understanding the ever-changing landscape of variants
Discovered on November 25 and designated a “variant of concern” by the WHO the following day, the Omicron variant (B.1.1.529) is being closely monitored by numerous research teams. Researchers from the “Infectious Diseases and Vectors: Ecology, Genetics, Evolution, and Control ” (University of Montpellier, CNRS, IRD), Mircea Sofonea, associate professor, and Samuel Alizon, research director, specialists in the epidemiology and evolution of infectious diseases, discuss the dynamics of variants. The prevalence of Delta, the characteristics of Omicron… Explanations in 10 key points by the two specialists.
Samuel Alizon, French National Research Institute for Sustainable Development (IRD) and Mircea T. Sofonea, University of Montpellier
The Conversation: Why did the Delta variant dominate all other SARS-CoV-2 variants for months?
Samuel Alizon: The Delta variant is quite “monstrous.” This is evident, for example, in estimates of the basic reproduction number, R₀ (the average number of infections caused by an infected person in a given population). Our team estimated it to be around 3 in a March 2020 report for ancestral lineages in France. For the Alpha variant, R₀ was between 4 and 5, which explains its rapid spread in early 2021. For Delta, estimates range from 6 to 8.
On pourrait presque parler d’un avantage « qualitatif » sur les autres variants, comme le signalent les études de terrain : si contrôler la propagation des lignées ancestrales revenait à stopper la propagation d’une grippe pandémique (R₀ < 3), avec Delta cela s’apparente plus à contrôler un virus comme la rubéole (R₀ > 5). Ce choc est particulièrement violent pour les populations peu vaccinées ou immunisées, comme on l’a vu cet été aux Antilles ou plus récemment en Europe de l’Est.
Mircea T. Sofonea: Among human respiratory viruses, only those that cause mumps, chickenpox, and measles are more contagious, with R₀ values often estimated at over 10. And the faster a virus spreads, the longer it will take for another variant—even one that is only slightly more contagious—to emerge.
Fisher’s geometric model, which dates back to the 1930s, illustrates a finding from evolutionary biology. In simple terms, it treats each mutation as a random step-by-step movement across a landscape whose topography represents the virus’s ability to spread within the human population. Through natural selection, only the steps that lead to an ascent are retained. Fisher’s model suggests that this ascent occurs increasingly slowly, as the probability that a random movement will land on a higher point decreases as it approaches the peak of the landscape—the adaptive optimum.
In fact, since July 2021, the number of variants has not skyrocketed; rather, we have seen the Delta variant lineage diversify into about a hundred sublineages —some of which are being monitored more closely due to the mutations of concern they carry.
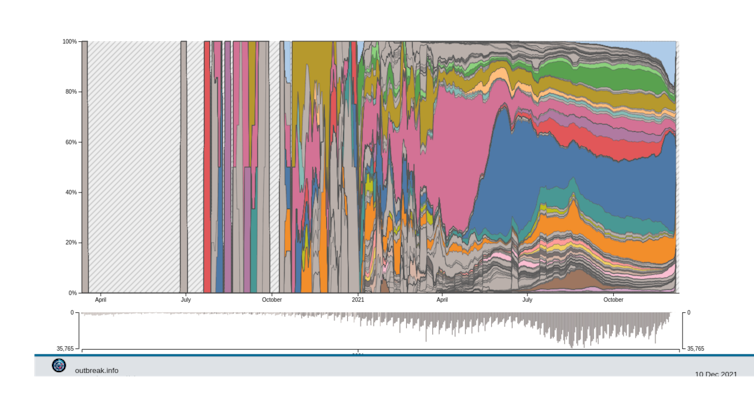
Delta Variant Report. Alaa Abdel Latif, Julia L. Mullen, Manar Alkuzweny, Ginger Tsueng, Marco Cano, Emily Haag, Jerry Zhou, Mark Zeller, Emory Hufbauer, Nate Matteson, Chunlei Wu, Kristian G. Andersen, Andrew I. Su, Karthik Gangavarapu, Laura D. Hughes
T.C.: Can Delta maintain this advantage “indefinitely”?
S.A.: The more the population becomes immune—whether through vaccination or, unfortunately, through infection itself—the more Delta’s advantage diminishes. Indeed, we now know that other variants are better at evading immunity than Delta. The most common hypothesis is therefore that Delta will ultimately be replaced by lineages capable of infecting immune hosts. For now, the Beta variant is the one for which laboratory tests detect the most immune escape.
Experiments using synthetic viral proteins make it possible to identify which mutations—or combinations of mutations—are most likely to emerge.
T.C.: You mentioned earlier that “we need to understand just how well the Delta variant has now adapted to us.” What is the situation?
S.A.: With a transmission rate twice that of the original strains, the Delta variant is undoubtedly well-adapted to our species in the short term. In the long term, it’s less certain: it will depend on the duration of our immunity against the infection, and the costs associated with immune escape for the virus. Indeed, we know that certain mutations allow the virus to evade the antibodies of recovered patients or vaccinated individuals… but we don’t know how contagious these mutated viruses are.
M.T.S.: It is important to keep in mind that the concept of adaptation, particularly in the case of an emerging viral disease, is relative: the adaptive landscape mentioned earlier is in fact characterized by movements comparable to ocean swells. The evolution of SARS-CoV-2 clearly illustrates the “arms race” we are engaged in with it: for the moment, we have, despite ourselves, selected more contagious and more virulent phenotypes (Alpha and Delta variants). Attention is now turning to variants capable of bypassing our second protective barrier: immunity (both post-vaccination and post-infection).
T.C.: Where might a new variant come from?
M.T.S.: A variant emerges just like any other mutation—randomly. Each of the nearly 30,000 bases (letters) in the SARS-CoV-2 genome mutates on average every 300,000 cell division cycles, and a single infection can produce several billion viral particles. Ultimately, the vast majority of infected individuals can transmit viruses that differ from those that infected them. It is estimated that, on average, along a chain of transmission, two mutations become fixed in the SARS-CoV-2 genome each month.
A particular mutant is considered a variant if it exhibits notable changes in one or more characteristics of interest (transmissibility, virulence, immune evasion, symptomatology, or antiviral resistance). The emergence of a variant often corresponds to a mutational surge, with evolutionary rates 2 to 4 times higher.
Ultimately, every infection that goes unchecked provides an opportunity for the virus to mutate and, potentially, give rise to a variant. Fortunately, such events remain very rare, as the majority of mutations are harmful.
It is the population in which the virus circulates that determines which mutations will be advantageous to it in that particular context (we say that selection pressures differ): if that population has no immunity at all, the most contagious strains are favored; if it is immune, then the strains capable of evading that immunity spread more widely.
S.A.: To this, we can add the case of chronic infections, particularly in immunocompromised individuals. In such cases, “intra-patient” selection occurs in addition to population-level selection. Indeed, it has been shown that during an infection lasting several months, the immune system selects SARS-CoV-2 viruses carrying mutations that have been found in variants.
In theory, this outcome is not automatic, and in the case of the human immunodeficiency virus (HIV), for example, it is believed that intra-patient adaptation occurs at the expense of transmission within the population. In any case, this finding means that the co-circulation of HIV and SARS-CoV-2 in unvaccinated populations, such as in sub-Saharan Africa, is a major public health concern, as was already noted during the emergence of the Gamma variant.
T.C.: What can we say more specifically about the origin of Omicron?
S.A.: The new Omicron variant was identified in South Africa, but it did not originate there. The country detected it thanks to the quality of its epidemiological and genomic surveillance. In this regard, the international community’s negative reaction toward South Africa is problematic, as it risks discouraging surveillance efforts.
While the exact origin of this variant is currently unknown, it most likely originated in a region of Africa where surveillance of the outbreak is limited. In fact, there are virtually no recent SARS-CoV-2 virus sequences similar to that of Omicron: genome analyses indicate that its common ancestor with the other variants dates back to mid-2020! This means it likely originated from lineages that circulated for over a year without being sampled (which is highly likely given the limited resources allocated to epidemic surveillance in many African countries).
It is also possible that the virus passed through an animal reservoir, as some of its mutations are puzzling. We know that SARS-CoV-2 can infect mammals, and in some cases—such as on mink farms—there have been instances of the virus spreading back to the human population. But for Omicron, there is still very little data available to explore this hypothesis.
T.C.: The term “evolutionary leap” has been used to describe certain variants. What does that mean?
S.A.: This touches on a long-standing debate in evolutionary biology between proponents of “gradualism”—a concept traceable back to Charles Darwin—and those of “punctuated equilibrium” (or “jumps”), popularized by Niles Eldredge and Stephen Jay Gould. The truth lies somewhere between the two, and the current pandemic provides a good example of this.
We know that SARS-CoV-2 naturally accumulates genetic mutations at a fairly steady rate, and this is what allows us to track it in phylodynamic studies. But the evolution of variants also results in widespread changes in the genome, or “selective sweeps” where a beneficial mutation (and associated sequences) becomes established: first with the D614G mutation (at the very start of the pandemic, the614th amino acid of the Spike protein was usually an aspartic acid— “D” in technical nomenclature; here it is replaced by glycine, “G,” ed.), followed by the Alpha variant (the Beta and Gamma variants remained in the minority globally), and then the Delta variant.
So depending on the time frame you’re looking at (a month or a year) and the criteria you use (neutral mutations or so-called phenotypic mutations that affect biological properties such as transmissibility), you’ll see either a continuous trend or sudden changes.
T.C.: And how should we interpret Omicron, which has 53 mutations, including about 30 in the spike protein alone?
S.A.: At this point, we know very little about Omicron, but its mutation profile is indeed striking. As South African viral evolution expert Darren Martin and his colleagues have analyzed, these mutations can be classified into three groups:
- On the one hand, there are all the mutations and deletions associated with the N501Y mutation in the spike protein (Δ69-70, K417N, N501Y, H655Y, P681H). The latter has already been described as profoundly altering the virus’s adaptive landscape—that is, the range of its possibilities.
- Next, still within the Spike protein, there is a second set of mutations at positions that have already been mutated in other variants, such as position 484 (N440S, S477I, or E484A). These are expected to have an effect on the phenotype of infections, such as their transmissibility, virulence, or ability to evade the immune response.
- Finally, there is a set of 14 mutations that are very rare in circulating lineages and virtually absent from other known sarbecoviruses (a subgenus of betacoronaviruses that includes coronaviruses associated with severe acute respiratory syndrome, such as SARS-CoV-2, ed.). Furthermore, individually, these mutations appear to be subject to negative selection. Their presence is therefore currently a mystery, as the variant appears well-adapted to our species. One possibility is that there is a collective effect of these mutations or an interaction with other mutations (such as N501Y) through a phenomenon of epistasis, common in population genetics: even if two mutations, A and B, are deleterious in isolation, their combined presence may be advantageous.
It should be noted that two sublineages of Omicron have already been identified. They are designated BA.1 and BA.2, and do not share the same key mutations.
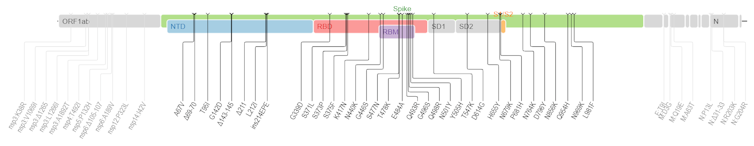
Mullen J, Tsueng G, et al., and tCfVS/https://covdb.stanford.edu/page/mutation-viewer/#omicron/Wikimedia, CC BY-SA
T.C.: What other signs would be cause for concern, aside from these mutations?
M.T.S.: The most striking point is that Omicron is linked to a very sharp resurgence of the epidemic in South Africa, immediately following the Delta variant wave. In this regard, it’s important to note that it’s very difficult to fully grasp national contexts. For example, our team is working hard to understand the epidemic in France. Consequently, to assess the severity of the epidemic situation in a country, it is best to rely on that country’s epidemiologists and health agencies… And in the case of South Africa, these specialists are concerned.
Several experiments are currently underway in which the virus is exposed to antibodies from vaccinated or recovered individuals. None of these results have been published yet, so it is difficult to draw conclusions, especially since preliminary results vary considerably from one study to another—and even within some studies…
It is important to keep in mind that these experiments indicate a general trend and do not capture the full diversity of the immune response. Ultimately, statistical analyses of epidemiological studies will be the most useful. In this regard, an initial field study suggests that the virus may cause more reinfections than other lineages. In other words, Omicron’s rapid spread may be explained more by its ability to evade immune responses than by its R₀.
T.C.: What might the actual global spread of Omicron be?
S.A.: Omicron has already been detected in small numbers in dozens of countries, including France. This pattern is reminiscent of what is known as a “source-sink” dynamic in ecological science: there would be a region of the world where this virus is predominant, and a process of dispersal is underway.
In several countries that are effectively tracking their outbreaks—notably the United Kingdom and Denmark—there is already a very rapid increase in test results consistent with this variant. Indeed, because it has a deletion at positions 69–70 of the spike protein, one of the screening tests—which targets three regions of the genome—yields a positive result if two of the three targets are present.
M.T.S.: In France, screening data—which involves looking for specific mutations—provides a near-real-time picture of how a group of genotypes is spreading. The advantage is that this technique is less expensive and faster than whole-genome sequencing. However, only whole-genome sequencing allows the variant to be identified with certainty.
T.C.: Do we have any idea how many variants (of interest or concern) may have emerged without being detected?
S.A.: All eyes are obviously on the new Omicron variant right now, but it’s hard to say exactly how many variants of concern exist worldwide. It’s quite likely that several have emerged without ever gaining traction. Indeed, even with a selective advantage, the early stages of a variant’s spread are governed by chance.
M.T.S.: In a simplified model, the probability of an epidemic dying out is 1/R₀. As a rough estimate, with an R₀ of 3, transmission chains die out spontaneously in 33% of cases. Thus, SARS-CoV-2 may have been introduced into France on multiple occasions, and the search for an index case is questionable.
This is also why it is possible that there have been several outbreaks of variants whose chains of transmission have died out on their own. Indeed, super-spreader events (such as gatherings without safety precautions, etc.) play a key role in the spread of this virus, with the vast majority of transmissions stemming from a small number of cases.
T.C.: What are the other variants currently under the closest scrutiny?
S.A.: In France, we are closely monitoring the B.1.640 lineage, which was first detected in March 2021, particularly in the Democratic Republic of the Congo; it is not classified as a variant of interest by the WHO but appears to be spreading fairly quickly. Our team has also identified the circulation of Delta variants carrying at least two mutations associated with immune escape in the spike protein (T95I and E484Q). For now, their circulation remains limited.
In any case, the emergence of the Omicron variant reminds us of the need for a long-term vision to overcome this pandemic. Apart from the scientific advisory board, few are concerned about this, as it conflicts with the immediacy of the political and media landscape. While it is just one player among many, scientific research has an important role to play, from developing treatments to anticipating the virus’s evolution.
Unfortunately, it is a long-term process, which makes it unappealing to policymakers… and it suffers from a lack of scientific literacy that pervades nearly every sector of French society. With the recent elimination of biology and physics-chemistry classes for most high school students, belief in miracle cures is likely to grow even stronger.![]()
Samuel Alizon, Director of Research CNRS, French National Research Institute for Sustainable Development (IRD) and Mircea T. Sofonea, Associate Professor of Epidemiology and the Evolution of Infectious Diseases, MIVEGEC Laboratory, University of Montpellier
This article is republished from The Conversation under a Creative Commons license. Readthe original article.