Phylodynamics: a “virus profiler”
An article in *Le Monde* published on April 22 provides further insight into a relatively new but particularly valuable field in COVID-19 research: phylodynamics. Among those cited is Samuel Alizon, a researcher at the University of Montpellier.
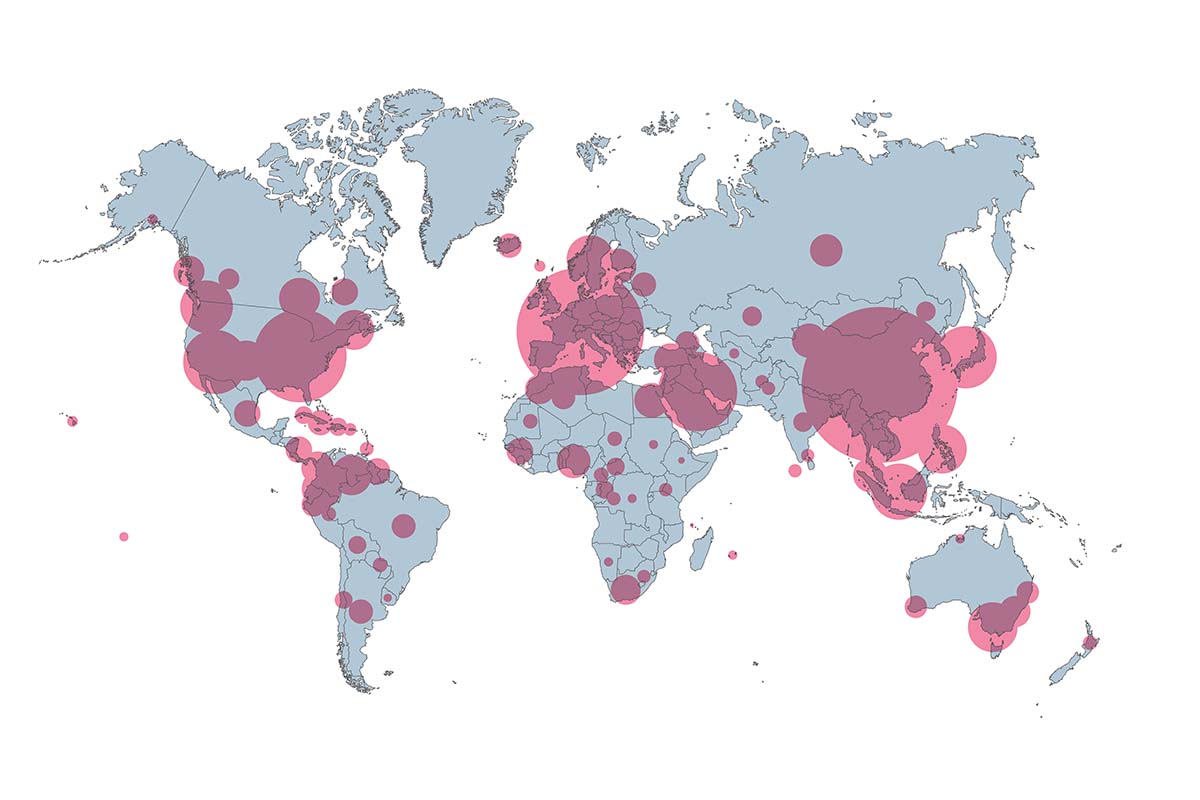
“The idea behind phylodynamics is that the way viruses spread leaves traces in their genomes,” explains Samuel Alizon, a researcher at the University of Montpellier in the Infectious Diseases and Vectors: Ecology, Genetics, Evolution, and Control (Mivegec) laboratory, in an article titled:“Phylodynamics: The Other Way to Track the Coronavirus,”published in Le Monde on April 22. These are minute traces, which the community of phylodynamics experts tracks by analyzing the thousands of DNA sequences of the virus now available.
The goal? To answer the countless questions raised by this pandemic. Where did this virus come from? When did it first infect humans? How fast is it spreading? How many people are potentially affected? These are all essential answers for a better understanding of the virus and, consequently, for anticipating its evolution—answers that the Montpellier-based researcher is analyzing with the help of mathematicians from LIRMM, in particular, and the PhyML software developed by Vincent Guindon, more specifically (read: PhyML, Montpellier-based software to trace the Covid-19 epidemic)
A fascinating and insightful article in *Le Monde*, in which internationally renowned scientists help us better understand the significance of this field, which didn’t even exist 20 years ago. The article also highlights the necessary precautions that must be taken when analyzing data from this highly nuanced science.